Genetic operators
Biological simulations and genetic algorithms alike use genetic operators to achieve its goals. These operators are potential bottlenecks of the programs/simulations, and given the size of the problems we want to address in the lab require for the highest optimization and efficiency, this work is used in several models.
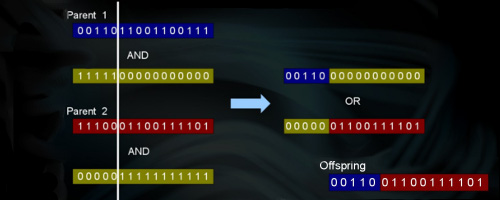
Several crossover (recombination) operators were optimized including the well known one-point, two-points, uniform, and half uniform. In addition, a skipping mechanism for fast processing of mutation was developed which works by ignoring a certain number of alleles in the genotype and only mutate those alleles that need to. Also, the traditional bit-flip mutation operator and the biologically realistic step mutation were sped-up.
The core part of this work lies in a versatile random number generator that allows for arbitrarily distributed pseudo-random numbers to be generated in only about three times the time that is required to generate an uniformly distributed random number.
Statistical tests are performed and C code is provided for ease of usage. The main intention of this work is to provide with a standard, ready to use, library of genetic operators for biological simulations and genetic algorithms.
The draft, tentatively called An efficient implementation of crossover and mutation in binary coded GAs, is still in preparation. We are currently thinking about submission to the Journal of Computational Biology. In addition, this work will be part of my Ph. D. dissertation.
About Me
![]() Edgar A. Duéñez Guzmán is a Senior Research Engineer at DeepMind.
Previously he was at Google, where he developed the first
machine learning system to select the index for Image Search.
During his academic career, he was a Postdoctoral fellow at the
Department of Biology at
KU Leuven working with
Tom Wenseleers in social evolution in microbes;
Edgar A. Duéñez Guzmán is a Senior Research Engineer at DeepMind.
Previously he was at Google, where he developed the first
machine learning system to select the index for Image Search.
During his academic career, he was a Postdoctoral fellow at the
Department of Biology at
KU Leuven working with
Tom Wenseleers in social evolution in microbes;
and a Research Associate at the
Department of Organismic and Evolutionary Biology at
Harvard University working with
David Haig in social evolution and imprinting.
Learn more...
Contact Info
E-mail: eaduenez {at} gmail {dot} com
 KU Leuven
KU Leuven OEB - Harvard
OEB - Harvard EECS - UTK
EECS - UTK FIT - Monash
FIT - Monash NIH
NIH HHMI
HHMI CIMAT
CIMAT


